This is a lovely tutorial from Torsten Seemann and team that demonstrates the use of GVL’s Galaxy server to do the common steps of bacterial genome analysis: assembly, variant calling and annotation.
The tutorial is here:
http://sepsis-omics.github.io/tutorials/
Before you get started, you need to add the tutorial dataset into your Galaxy instance. Download it from http://sepsis-omics.github.io/tutorials/modules/data-dna/Data.tar.gz and extract it to a local directory.
Use Galaxy to upload the data.
There is also a quick fix you need to do to make Prokka work (will be fixed in next GVL release), so please login to your server with SSH and execute the following two commands:
cd /mnt/gvl
sudo gvl/apps/linuxbrew/bin/* ./galaxy/tools/hmmer/3.1b2/iuc/package_hmmer_3_1b2/2a1881410dc9/bin/
First, if you haven’t already, use the Bryn interface to start your GVL server.
Follow the link to your server to the GVL homepage:

Click the link to Galaxy:
Sign-up for a Galaxy account using any details you wish.
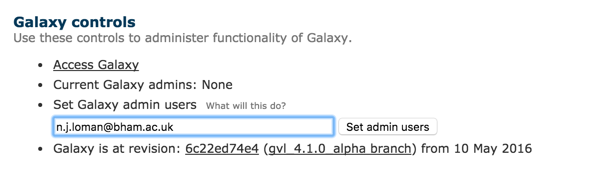
Then you need to make yourself an administrator. Go to the Admin page on your GVL instance and add the email address you signed up with and choose ‘Set admin users’.
Now follow the tutorial over at: